This article demonstrates the two multi-horizon superior predictive
ability tests from Quaedvlieg (2021), implemented in
forecastdom as uspa_mh_test() and
aspa_mh_test(). Both tests compare a path of
forecasts at horizons
rather than each horizon in isolation, and use a moving-block bootstrap
to obtain critical values that account for serial dependence in the
loss-differential path.
- Uniform SPA () — null: the benchmark is at least as good as the competitor at every horizon. Test statistic is the minimum of horizon-wise standardized loss differentials.
- Average SPA () — null: the benchmark is at least as good as the competitor on a (user-specified) weighted average of horizons. Test statistic is the standardized weighted average loss differential.
Throughout, the loss differential is defined as , so positive entries indicate the benchmark has higher loss (is worse) at that horizon, and rejection of either null requires the benchmark to be uniformly (or on average) worse.
library(forecastdom)
data(quaedvlieg2021)
str(quaedvlieg2021, max.level = 1)
#> List of 2
#> $ uspa: num [1:1000, 1:20] 1.189 -5.113 -1.075 -0.631 -3.759 ...
#> $ aspa: num [1:1000, 1:20] 1.171 -5.13 -1.092 -0.649 -3.776 ...The bundled quaedvlieg2021 object holds the two
loss-differential matrices distributed with the paper’s replication
archive: $uspa and $aspa, each
.
Per-horizon picture
Before running the tests, it’s worth looking at the per-horizon mean loss differentials. The two example matrices are constructed so that they discriminate between the tests.
H <- ncol(quaedvlieg2021$uspa)
means <- rbind(
uspa_dataset = colMeans(quaedvlieg2021$uspa),
aspa_dataset = colMeans(quaedvlieg2021$aspa)
)
colnames(means) <- paste0("h", seq_len(H))
knitr::kable(round(means, 3))| h1 | h2 | h3 | h4 | h5 | h6 | h7 | h8 | h9 | h10 | h11 | h12 | h13 | h14 | h15 | h16 | h17 | h18 | h19 | h20 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| uspa_dataset | 0.004 | 0.136 | 0.182 | 0.164 | 0.135 | 0.092 | 0.185 | 0.224 | 0.260 | 0.171 | 0.272 | 0.280 | 0.262 | 0.232 | 0.274 | 0.332 | 0.383 | 0.351 | 0.411 | 0.425 |
| aspa_dataset | -0.014 | 0.101 | 0.140 | 0.116 | 0.082 | 0.035 | 0.124 | 0.160 | 0.193 | 0.101 | 0.199 | 0.204 | 0.184 | 0.151 | 0.191 | 0.247 | 0.295 | 0.261 | 0.319 | 0.330 |
$uspa shows positive mean loss differentials at
every horizon — the benchmark is worse uniformly.
$aspa shows positive means at most horizons but a near-zero
mean at the shortest horizon, so the benchmark is worse on average but
not uniformly.
library(ggplot2)
df <- data.frame(
horizon = rep(seq_len(H), 2),
d_bar = c(colMeans(quaedvlieg2021$uspa),
colMeans(quaedvlieg2021$aspa)),
dataset = rep(c("uspa", "aspa"), each = H)
)
ggplot(df, aes(horizon, d_bar, colour = dataset)) +
geom_hline(yintercept = 0, linetype = "dashed", colour = "grey60") +
geom_line() + geom_point() +
labs(x = "Horizon h", y = expression(bar(d)[h]),
title = "Mean loss differential by horizon") +
theme_minimal()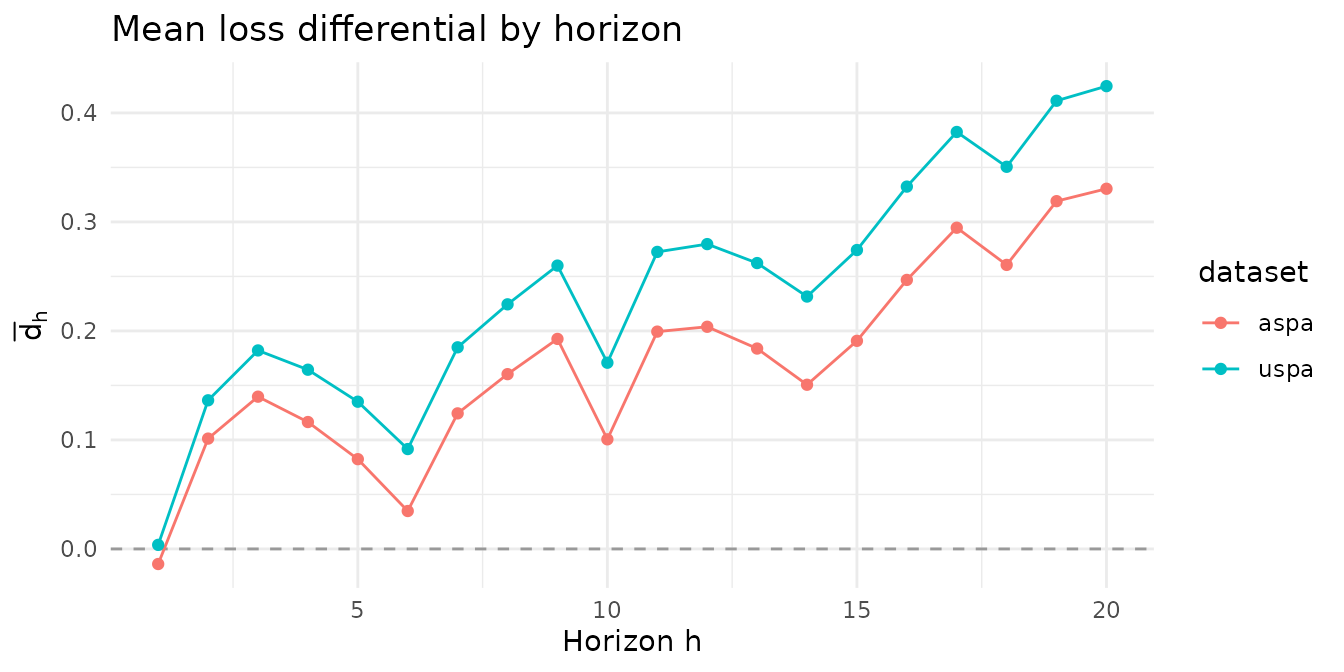
Uniform multi-horizon SPA
set.seed(1)
uspa_uspa <- uspa_mh_test(quaedvlieg2021$uspa, L = 3, B = 999)
set.seed(1)
uspa_aspa <- uspa_mh_test(quaedvlieg2021$aspa, L = 3, B = 999)
uspa_uspa
uspa_aspa
#>
#> ╭────────────────────────────────────────────────────╮
#> │ Uniform Multi-Horizon SPA Test │
#> │ (Quaedvlieg, 2021) │
#> ├────────────────────────────────────────────────────┤
#> │ H0: Benchmark has uSPA (weakly dominates) │
#> │ H1: Benchmark uniformly worse than competitor │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Test Results: │
#> │ uSPA statistic: 0.0804 │
#> │ P-value (MBB): 0.0240 │
#> │ Decision: Rejected *** │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Details: │
#> │ Observations (T): 1000 │
#> │ Horizons (H): 20 │
#> │ Block length (L): 3 │
#> │ Bootstrap replications: 999 │
#> │ Significance level: 0.0500 │
#> ╰────────────────────────────────────────────────────╯
#>
#>
#> ╭────────────────────────────────────────────────────╮
#> │ Uniform Multi-Horizon SPA Test │
#> │ (Quaedvlieg, 2021) │
#> ├────────────────────────────────────────────────────┤
#> │ H0: Benchmark has uSPA (weakly dominates) │
#> │ H1: Benchmark uniformly worse than competitor │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Test Results: │
#> │ uSPA statistic: -0.2980 │
#> │ P-value (MBB): 0.0751 │
#> │ Decision: Not rejected │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Details: │
#> │ Observations (T): 1000 │
#> │ Horizons (H): 20 │
#> │ Block length (L): 3 │
#> │ Bootstrap replications: 999 │
#> │ Significance level: 0.0500 │
#> ╰────────────────────────────────────────────────────╯The uSPA test rejects on $uspa (all horizons show the
benchmark losing) but fails to reject on $aspa at the 5%
level, because the shortest horizon’s mean is essentially zero — the
min of standardized horizon-wise means is pulled down by
that single horizon.
Average multi-horizon SPA
With uniform weights the aSPA statistic is the standardized unweighted average loss differential:
w_unif <- rep(1 / H, H)
set.seed(1)
aspa_uspa <- aspa_mh_test(quaedvlieg2021$uspa, weights = w_unif, L = 3, B = 999)
set.seed(1)
aspa_aspa <- aspa_mh_test(quaedvlieg2021$aspa, weights = w_unif, L = 3, B = 999)
aspa_uspa
aspa_aspa
#>
#> ╭────────────────────────────────────────────────────╮
#> │ Average Multi-Horizon SPA Test │
#> │ (Quaedvlieg, 2021) │
#> ├────────────────────────────────────────────────────┤
#> │ H0: Benchmark has aSPA (weighted avg superior) │
#> │ H1: Benchmark worse on weighted average │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Test Results: │
#> │ aSPA statistic: 3.4075 │
#> │ P-value (MBB): 0.0010 │
#> │ Decision: Rejected *** │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Details: │
#> │ Observations (T): 1000 │
#> │ Horizons (H): 20 │
#> │ Block length (L): 3 │
#> │ Bootstrap replications: 999 │
#> │ Significance level: 0.0500 │
#> ╰────────────────────────────────────────────────────╯
#>
#>
#> ╭────────────────────────────────────────────────────╮
#> │ Average Multi-Horizon SPA Test │
#> │ (Quaedvlieg, 2021) │
#> ├────────────────────────────────────────────────────┤
#> │ H0: Benchmark has aSPA (weighted avg superior) │
#> │ H1: Benchmark worse on weighted average │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Test Results: │
#> │ aSPA statistic: 2.4397 │
#> │ P-value (MBB): 0.0040 │
#> │ Decision: Rejected *** │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Details: │
#> │ Observations (T): 1000 │
#> │ Horizons (H): 20 │
#> │ Block length (L): 3 │
#> │ Bootstrap replications: 999 │
#> │ Significance level: 0.0500 │
#> ╰────────────────────────────────────────────────────╯The aSPA test rejects on both datasets — the negative h=1
mean of $aspa is more than compensated by positive means at
longer horizons, so the weighted average favours rejection. This is the
power gain Quaedvlieg (2021) emphasises: by aggregating across the
forecast path, the average test detects model-level differences that the
per-horizon Diebold-Mariano machinery would miss because of the
multiple-testing burden.
Custom weights
Down-weighting short horizons makes the aSPA test even more decisive
against $aspa:
w_down <- c(rep(0, 4), rep(1, 16)) / 16 # zero weight on h = 1..4
set.seed(1)
aspa_mh_test(quaedvlieg2021$aspa, weights = w_down, L = 3, B = 999)
#>
#> ╭────────────────────────────────────────────────────╮
#> │ Average Multi-Horizon SPA Test │
#> │ (Quaedvlieg, 2021) │
#> ├────────────────────────────────────────────────────┤
#> │ H0: Benchmark has aSPA (weighted avg superior) │
#> │ H1: Benchmark worse on weighted average │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Test Results: │
#> │ aSPA statistic: 2.3237 │
#> │ P-value (MBB): 0.0060 │
#> │ Decision: Rejected *** │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Details: │
#> │ Observations (T): 1000 │
#> │ Horizons (H): 20 │
#> │ Block length (L): 3 │
#> │ Bootstrap replications: 999 │
#> │ Significance level: 0.0500 │
#> ╰────────────────────────────────────────────────────╯Up-weighting the shortest horizons pulls the statistic back toward zero:
w_up <- c(rep(4, 4), rep(0, 16)) / 16 # all weight on h = 1..4
set.seed(1)
aspa_mh_test(quaedvlieg2021$aspa, weights = w_up, L = 3, B = 999)
#>
#> ╭────────────────────────────────────────────────────╮
#> │ Average Multi-Horizon SPA Test │
#> │ (Quaedvlieg, 2021) │
#> ├────────────────────────────────────────────────────┤
#> │ H0: Benchmark has aSPA (weighted avg superior) │
#> │ H1: Benchmark worse on weighted average │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Test Results: │
#> │ aSPA statistic: 1.9568 │
#> │ P-value (MBB): 0.0210 │
#> │ Decision: Rejected *** │
#> ├┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┄┤
#> │ Details: │
#> │ Observations (T): 1000 │
#> │ Horizons (H): 20 │
#> │ Block length (L): 3 │
#> │ Bootstrap replications: 999 │
#> │ Significance level: 0.0500 │
#> ╰────────────────────────────────────────────────────╯Block-length sensitivity
The moving-block bootstrap depends on the block length
L. The p-value is robust over a reasonable range — small
enough to keep many blocks, large enough to capture the path’s serial
dependence:
Ls <- c(2, 3, 5, 8, 12)
sens <- do.call(rbind, lapply(Ls, function(L) {
set.seed(1)
u <- uspa_mh_test(quaedvlieg2021$uspa, L = L, B = 499)
set.seed(1)
a <- aspa_mh_test(quaedvlieg2021$uspa, weights = w_unif, L = L, B = 499)
data.frame(L = L,
uspa_stat = u$statistic, uspa_p = u$pvalue,
aspa_stat = a$statistic, aspa_p = a$pvalue)
}))
knitr::kable(
sens, digits = 3, row.names = FALSE,
col.names = c("$L$",
"uSPA stat", "uSPA $p$",
"aSPA stat", "aSPA $p$"))| uSPA stat | uSPA | aSPA stat | aSPA | |
|---|---|---|---|---|
| 2 | 0.08 | 0.028 | 3.407 | 0.000 |
| 3 | 0.08 | 0.020 | 3.407 | 0.002 |
| 5 | 0.08 | 0.030 | 3.407 | 0.002 |
| 8 | 0.08 | 0.036 | 3.407 | 0.002 |
| 12 | 0.08 | 0.024 | 3.407 | 0.000 |
The statistics are independent of L (they depend only on
the QS HAC long-run variance, not on the bootstrap), and the p-values
are stable across L for this dataset.
Takeaways
-
uspa_mh_test()rejects only when the benchmark loses at every horizon — strong power against uniform underperformance, limited power against horizon-specific failures. -
aspa_mh_test()rejects when the weighted-average differential favours the competitor, even if performance at individual horizons is mixed. The choice of weights determines which segment of the forecast path drives the decision. - The moving-block bootstrap is essential: forecast paths exhibit natural serial dependence, especially when the same model is evaluated at successive horizons.
